| Door | Female | Male | Difference (M <U+2212> F) |
|---|---|---|---|
| Door A | 162.1 | 175.2 | 13.1 |
| Door B | 163.4 | 176.6 | 13.2 |
| Door C | 162.0 | 177.2 | 15.2 |
| Door D | 161.9 | 175.3 | 13.4 |
Missing Data
MCAR, MAR, and MNAR
Welcome, Players
You are data analysts at a dating app. Your company collected data on 1,000 users — height, sex, and whether they completed their full profile.
But something went wrong. The data was incomplete, and now four different versions exist. Only one is the original.
Your goal is to pick a door, analyze the data behind it, and figure out if your data has the truth.

Round 1: Pick a Door
The Four Doors
 :::
:::
Behind each door is a version of your dating app data with the same 1,000 users. Three were corrupted by a different unknown force. One was left untouched.
You must choose one door. That’s your dataset.
One door has no missing values at all, you know the truth, so please don’t say your data is complete.
Choose Your Door
Run the following code to randomly select your door:
Write down which door you got. Now let’s see what’s behind it.
Open Your Door
Compute the group means from your dataset:
Write down your results:
- Male mean height = ?
- Female mean height = ?
- How many missing values do you have?
We’ll compare answers across the class in a moment.
Look around the zoom:
- Did everyone get the same means?
- Did anyone have zero missing values? (don’t tell)
The force behind each door determined whether your results are trustworthy.
All Four Doors Opened
Let’s see what was behind every door:
Round 2: The Three Forces
Naming the Forces
Rubin [-@rubin1976inference] identified three forces that make data disappear. Each one leaves a different trace and requires a different strategy.
| Force (Full Name) | Missingness depends on… |
|---|---|
| MCAR (Missing Completely At Random) | Nothing — purely random |
| MAR (Missing At Random) | Observed variables only |
| MNAR (Missing Not At Random) | The missing values themselves |
Learn to recognize each one, then we’ll see which force hit your door.
Force 1: MCAR — The Random Storm
Definition: The probability of being missing is unrelated to any variable observed or unobserved.
\[P(R = 1 \mid \text{Sex}, \text{Height}) = P(R = 1)\]
In plain language: This force strikes like lightning completely at random. It doesn’t care who you are or what your height is.
Where you’ll encounter it:
- A server crash deletes random records
- You don’t ask for height to some users (randomly)
- People did not see the height question because it was behind a banner that appears randomly
Good news: Complete-case analysis is unbiased; you lose statistical power.
Planned Missing Data Designs (MCAR by Design)
Researchers sometimes intentionally introduce MCAR to reduce participant burden while preserving statistical power.
Three-form design [@Graham_2006]:
- All participants answer a common block (X)
- Each participant is randomly assigned two of three additional blocks (A, B, C)
- Every item pair appears together in at least one form
| Form | Blocks |
|---|---|
| 1 | X + A + B |
| 2 | X + A + C |
| 3 | X + B + C |
Why do this?
- Cuts survey length by ~33% per participant
- Reduces fatigue and careless responding
- Missingness is MCAR by construction (random assignment to forms)
- MI or FIML recovers full-data estimates with minimal efficiency loss
Other planned designs:
- Two-method measurement - expensive gold-standard measure on a random subsample, cheap proxy for everyone
- Wave-missing - in longitudinal studies, skip a random subset of participants at certain waves
Force 2: MAR - The Hidden Trail
Definition: The probability of being missing depends on observed data, but not on the missing value itself.
\[P(R = 1 \mid Y_{\text{obs}}, Y_{\text{mis}}) = P(R = 1 \mid Y_{\text{obs}})\]
In plain language: This force follows a trail you can see. If you find the trail, you can undo the damage.
Where you’ll encounter it:
- Users who skip their full profile also skip reporting height
- Newer users are less likely to fill in optional fields
- Users who never upload a photo also leave height blank
Key: Complete-case analysis can be biased, but proper methods (MI, FIML) can correct it if you follow the right trail.
Force 3: MNAR - The Invisible Wall
Definition: The probability of being missing depends on the missing value itself, even after controlling for observed data.
\[P(R = 1 \mid \text{Sex}, \text{Height}) \newline \text{depends directly on height}\]
In plain language: This is the most dangerous force. The data hides because of what it contains. There is no visible trail to follow.
Where you’ll encounter it:
- Shorter men don’t report their height on dating apps
- High earners refuse to report income
- Sickest patients drop out of clinical trials
Warning: No standard method can fully correct this bias. You’ll need sensitivity analysis.
Your Map of the Forces
| Feature | MCAR (Random Storm) | MAR (Hidden Trail) | MNAR (Invisible Wall) |
|---|---|---|---|
| Depends on | Nothing | Observed variables | The missing values |
| Complete-case bias? | No | Yes | Yes |
| Fixable with MI/FIML? | Yes (N/A) | Yes | Not fully |
| How common? | Rare | Common | Common |
| Testable? | Partially | Not definitively | Not definitively |
Important
We can never fully verify whether data is MAR or MNAR from the observed data alone. The classification requires substantive reasoning about the data-generating process.
Now, can you match each force to one of the doors?
Round 3: The Reveal
How It Works
I’m going to show you the code used to generate each door’s dataset.
Your job: read the code and tell me what mechanism it is.
- Is missingness related to nothing? → MCAR
- Is missingness related to an observed variable? → MAR
- Is missingness related to the missing value itself? → MNAR
Door A: The Code
What do you see?
- What determines whether height goes missing?
- Does it depend on sex? On height? On profile status?
MCAR — The Random Storm. Every observation has the same 40% chance of being missing. It depends on nothing — pure random chance. The complete cases are a random subset of the data. You lose power, but the truth survives.
Door B: The Code
First, how profile_status was generated:
Then missingness in height depends on profile_status:
What do you see?
profile_statusdepends on height; shorter users are less likely to complete their profile- Missingness in height depends on
profile_status; not on height directly
MAR - The Hidden Trail. Missingness depends on an observed variable (
profile_status). Since shorter users tend to have incomplete profiles (and thus more missing height), the naive group means are biased upward. If we control forprofile_status, we can recover the truth.
Door B: Controlling for Profile Status
Try it yourself — estimate the mean height by sex, controlling for profile status:
We can also recover the marginal means using regression + predict:
Door C: The Code
What do you see?
- What determines whether height goes missing for males?
- Can you observe
heightif it’s the variable that went missing?
MNAR - The Invisible Wall. Among males, missingness depends on height itself; the variable that’s missing. Shorter males have up to 60% chance of being missing. There’s no observed trail to follow.
No standard method can fully correct this.
Door D: The Code
No problem at all. Door D is the complete, original data; no missing values, no mechanism.
If you chose this door, you had the truth the whole time.
Ground truth: Male = 175.3, Female = 161.9
The Evidence Board
Now let’s compare all four doors against the truth:
| Door | Difference | Female | Male |
|---|---|---|---|
| Door D (Truth) | 13.4 | 161.9 | 175.3 |
| Door A (MCAR) | 13.1 | 162.1 | 175.2 |
| Door B (MAR) | 13.2 | 163.4 | 176.6 |
| Door C (MNAR) | 15.2 | 162.0 | 177.2 |
Door A (MCAR): close to the truth; random loss, no bias
Door B (MAR): biased; but fixable if we control for
profile_status
Door C (MNAR): biased; and not fixable with standard methods
Visualizing the Damage
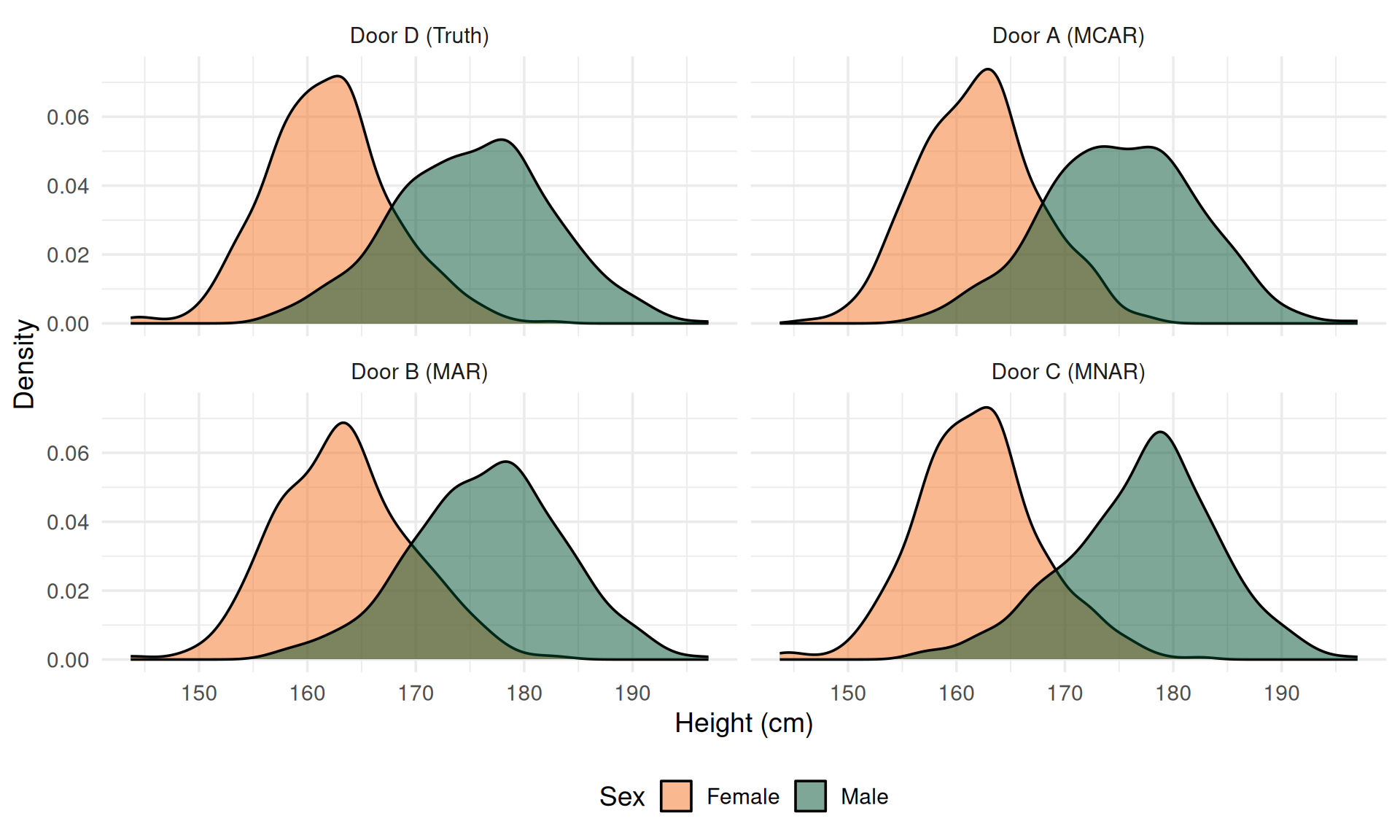
MAR: Why Is It Biased?
profile_status depends on height. Shorter users are less likely to complete their profile:
| sex | profile_status | n | mean_height |
|---|---|---|---|
| Male | Incomplete | 216 | 172.4 |
| Male | Complete | 284 | 177.5 |
| Female | Incomplete | 353 | 160.8 |
| Female | Complete | 147 | 164.6 |
Shorter users → incomplete profile → more missing height → observed means are biased upward.
MAR: Following the Trail
But within each profile-status group, missing and observed cases have similar mean heights:
| profile_status | status | n | mean_height |
|---|---|---|---|
| Incomplete | Missing | 402 | 164.7 |
| Incomplete | Observed | 167 | 166.5 |
| Complete | Observed | 402 | 173.0 |
| Complete | Missing | 29 | 174.5 |
This is the signature of MAR. If we include profile_status as an auxiliary variable in MI/FIML, we recover unbiased estimates.
MNAR: Behind the Wall
| profile_status | status | n | mean_height |
|---|---|---|---|
| Incomplete | Observed | 455 | 164.9 |
| Incomplete | Missing | 114 | 166.4 |
| Complete | Observed | 345 | 173.5 |
| Complete | Missing | 86 | 171.6 |
Among males, missing cases are shorter than observed cases. The data disappears because of its own value. No observed variable can explain this; the bias is not fixable with standard methods.
Mapping the Three Forces
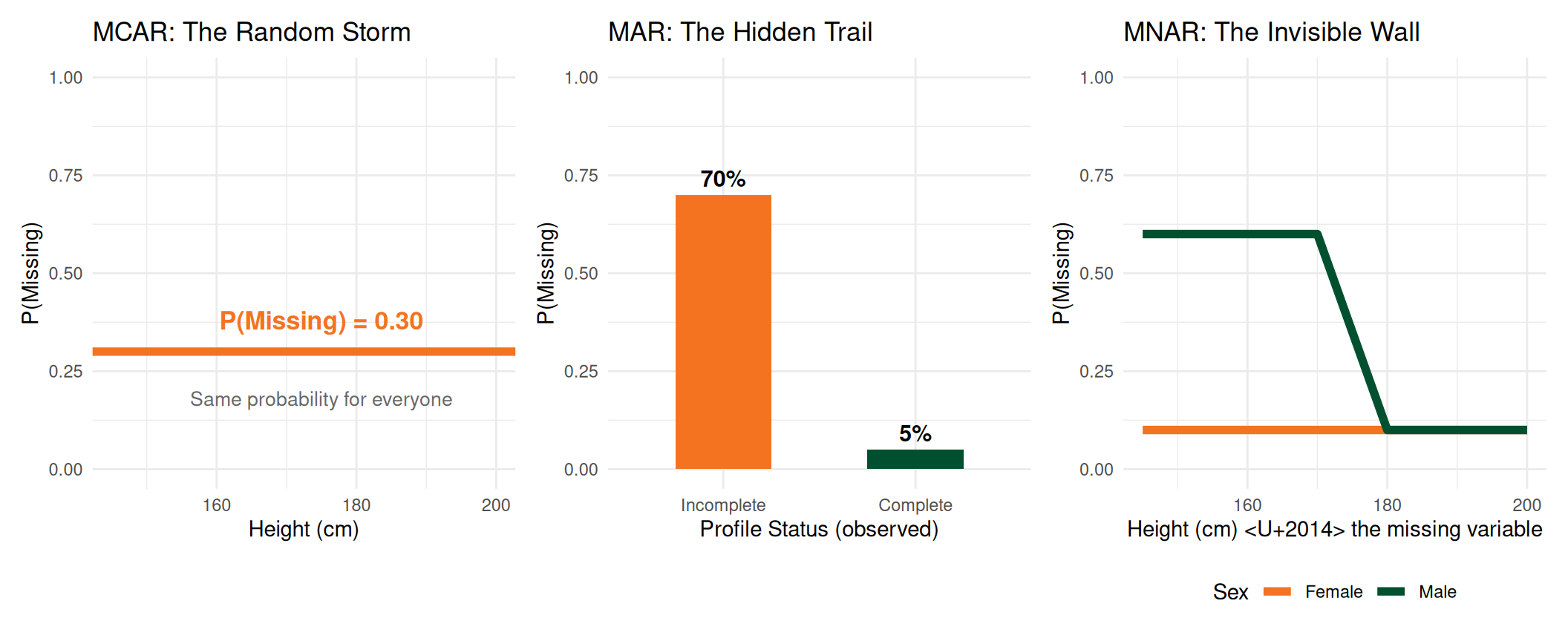
The Takeaway
If you ignore the forces:
- Group means are biased
- Treatment effects are distorted
- Confidence intervals miss the truth
- p-values are unreliable
- Effect sizes are attenuated
Real-world consequences:
- Clinical trials underestimate side effects
- Surveys underrepresent vulnerable groups
- Longitudinal studies suffer from survivor bias
- Policy decisions rest on distorted evidence
The method for handling missing data should be informed by the mechanism that generated it. Rubin
Apply to Your Own Research
Consider your own research or a study you’ve recently read:
- Which variables might have missing data?
- What force (MCAR, MAR, MNAR) is most plausible? Why?
- What would happen if you just deleted all incomplete cases?
- What observed variables might predict the missingness?
Lessons Learned
- Missing data mechanisms matter. They determine whether your results are biased
- MCAR (the random storm) is the only safe scenario for complete-case analysis and it’s the rarest
- MAR (the hidden trail) can be addressed with proper methods if you follow the right auxiliary variables
- MNAR (the invisible wall) is the hardest challenge requires substantive reasoning and sensitivity analysis
- Always ask why data is missing, not just how much
Part II: The Toolkit
Equipping Yourself
Now that you can name the forces, you need the right tools to deal with them.
- Complete-case analysis: Deleting incomplete cases
- Multiple Imputation (MI): Creating plausible completed datasets
- Full Information Maximum Likelihood (FIML): Using all available information
- Sensitivity analysis: What to do when you suspect MNAR
Multiple Imputation: The Big Idea
Instead of filling in one guess for each missing value, MI creates m plausible completed datasets.
Three steps:
- Impute: Generate m completed datasets
- Analyze: Fit your model on each one
- Pool: Combine with Rubin’s Rules
Each imputed dataset is slightly different, reflecting our uncertainty about what the missing values could have been.
Pooled estimate:
\[\bar{Q} = \frac{1}{m} \sum_{i=1}^{m} \hat{Q}_i\]
Total variance:
\[T = \bar{U} + \left(1 + \frac{1}{m}\right) B\]
Where \(\bar{U}\) = within-imputation variance and \(B\) = between-imputation variance.
MICE: Chained Equations
MICE = Multivariate Imputation by Chained Equations
Each variable with missing data gets its own imputation model, using all other variables as predictors.
The algorithm:
- Fill in starting values (e.g., random draws from observed)
- For each variable with missing data:
- Regress it on all other variables (observed cases)
- Draw imputed values from the predictive distribution
- Repeat step 2 for many iterations (cycling)
- Store the final imputed dataset
- Repeat m times → m complete datasets
Default methods in mice:
| Variable Type | Method |
|---|---|
| Continuous | Predictive mean matching |
| Binary (2 lev.) | Logistic regression |
| Nominal (>2 lev.) | Multinomial logit |
| Ordinal | Proportional odds |
A Simulated Clinical Trial
We simulate a randomized trial with 200 participants, two binary predictors, three continuous, and one auxiliary variable (stress):
set.seed(752)
n_mi <- 200
mi_complete <- tibble(
id = 1:n_mi,
age = round(rnorm(n_mi, mean = 0, sd = 1), 2),
female = rbinom(n_mi, 1, 0.55),
treatment = rbinom(n_mi, 1, 0.5),
baseline_severity = round(rnorm(n_mi, mean = 0, sd = 1), 2),
social_support = round(rnorm(n_mi, mean = 0, sd = 1), 2),
stress = round(rnorm(n_mi, mean = 0, sd = 1), 2),
outcome = round(
-0.5 * treatment - 0.3 * age + 0.4 * female +
0.5 * baseline_severity - 0.3 * social_support +
0.8 * stress + rnorm(n_mi, 0, 1), 2
)
)stress is an auxiliary variable: it affects the outcome but will not be included in the analysis model. It will, however, drive missingness and be available to MICE.
True parameters (analysis model): \(\beta_{\text{trt}} = -0.5\), \(\beta_{\text{age}} = -0.3\), \(\beta_{\text{female}} = 0.4\), \(\beta_{\text{severity}} = 0.5\), \(\beta_{\text{support}} = -0.3\)
Making Data Disappear (MAR)
We corrupt the data with MAR missingness. The probability of being missing depends on observed variables:
mi_data <- mi_complete |>
mutate(
# stress (auxiliary) + treatment drive outcome missingness
p_miss_out = plogis(-0.5 + 1.5 * stress + 1.0 * treatment),
# Higher age -> more likely to miss social support
p_miss_ss = plogis(-0.5 + 1.2 * age),
outcome = ifelse(rbinom(n(), 1, p_miss_out) == 1, NA, outcome),
social_support = ifelse(rbinom(n(), 1, p_miss_ss) == 1, NA, social_support)
) |>
select(-p_miss_out, -p_miss_ss)This is MAR because missingness depends only on fully observed variables — stress and treatment drive missingness in outcome, while age drives missingness in social_support. The key twist: stress affects the outcome (\(\beta = 0.8\)) but is not in the analysis model. Complete-case analysis is biased because dropping high-stress treated participants distorts the treatment–outcome relationship (collider bias: treatment \(\rightarrow\) R \(\leftarrow\) stress). MI recovers because stress is included in the imputation model as an auxiliary variable.
Inspect the Damage
| Variable | N_Missing | Pct_Missing |
|---|---|---|
| social_support | 95 | 38 |
| outcome | 125 | 50 |
age female treatment baseline_severity stress social_support outcome
78 1 1 1 1 1 1 1 0
77 1 1 1 1 1 1 0 1
47 1 1 1 1 1 0 1 1
48 1 1 1 1 1 0 0 2
0 0 0 0 0 95 125 220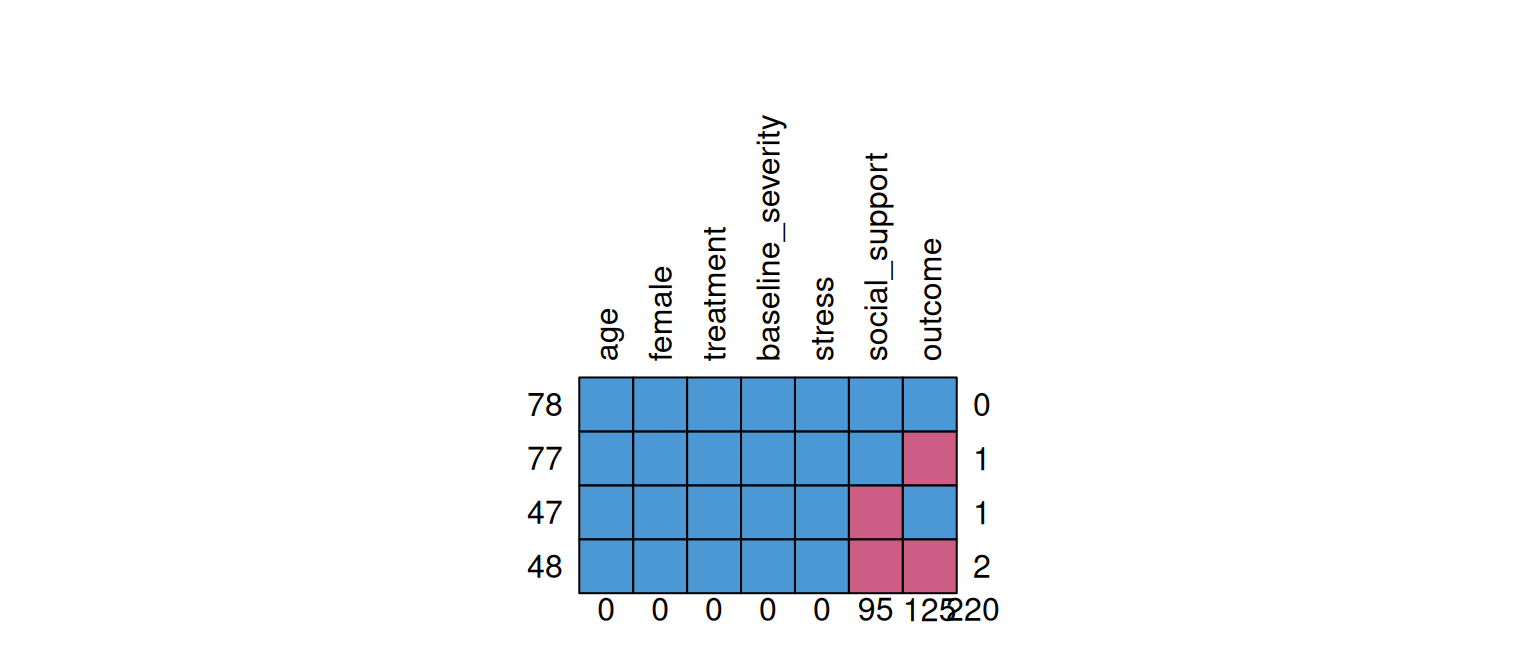
Each row is a pattern (1 = observed, 0 = missing). The numbers show how many cases share each pattern.
Step 1: Run MICE
Class: mids
Number of multiple imputations: 5
Imputation methods:
age female treatment baseline_severity
"" "" "" ""
social_support stress outcome
"pmm" "" "pmm"
PredictorMatrix:
age female treatment baseline_severity social_support stress
age 0 1 1 1 1 1
female 1 0 1 1 1 1
treatment 1 1 0 1 1 1
baseline_severity 1 1 1 0 1 1
social_support 1 1 1 1 0 1
stress 1 1 1 1 1 0
outcome
age 1
female 1
treatment 1
baseline_severity 1
social_support 1
stress 1mice automatically selects predictive mean matching (pmm) for numeric variables. All fully observed variables serve as predictors in each imputation model.
Step 2: Check Convergence
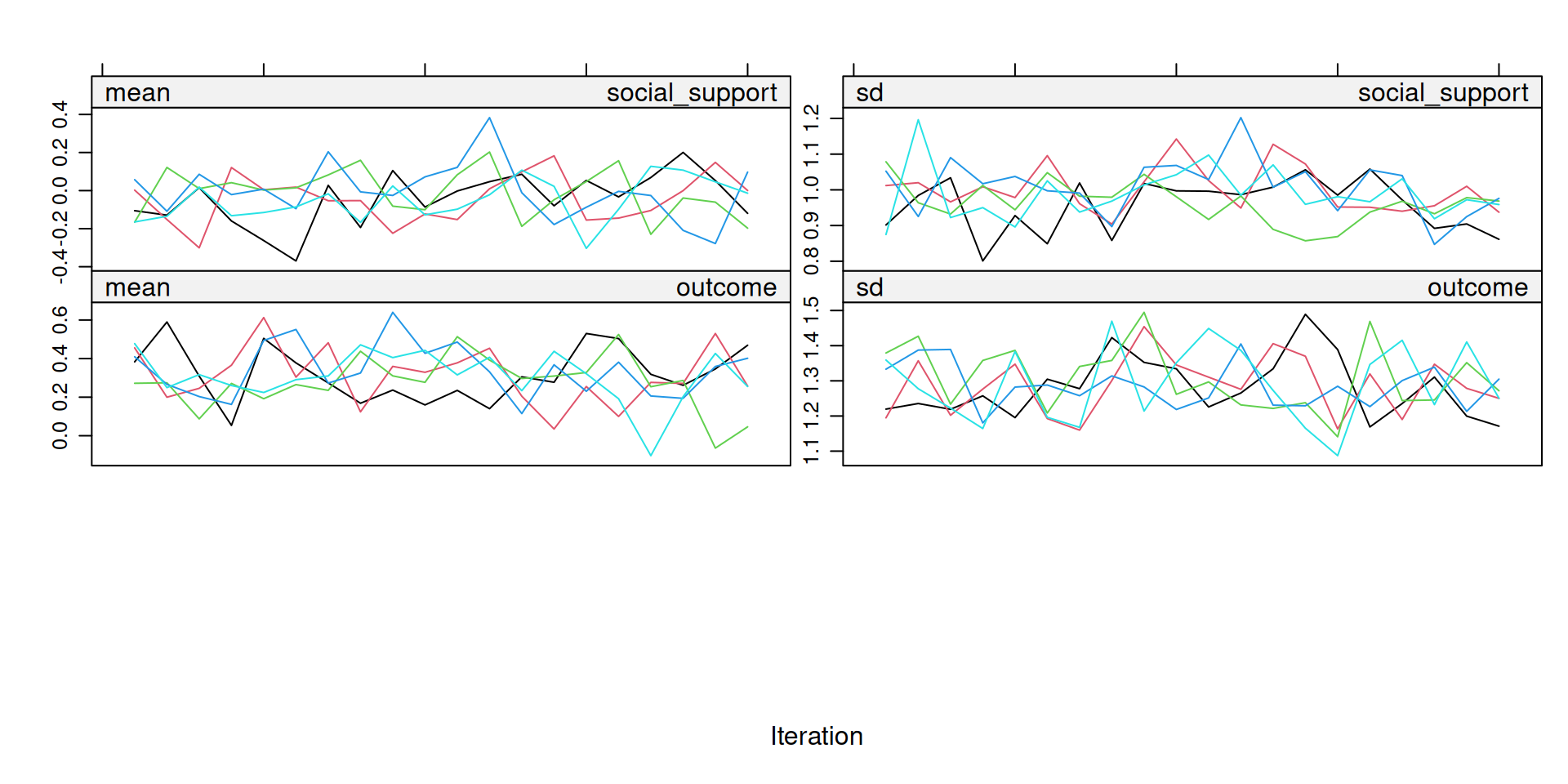
What to look for: The m colored streams should be freely intermixed with no systematic trend. If streams separate or drift upward/downward check the imputation model.
Step 3: Inspect Imputed Values
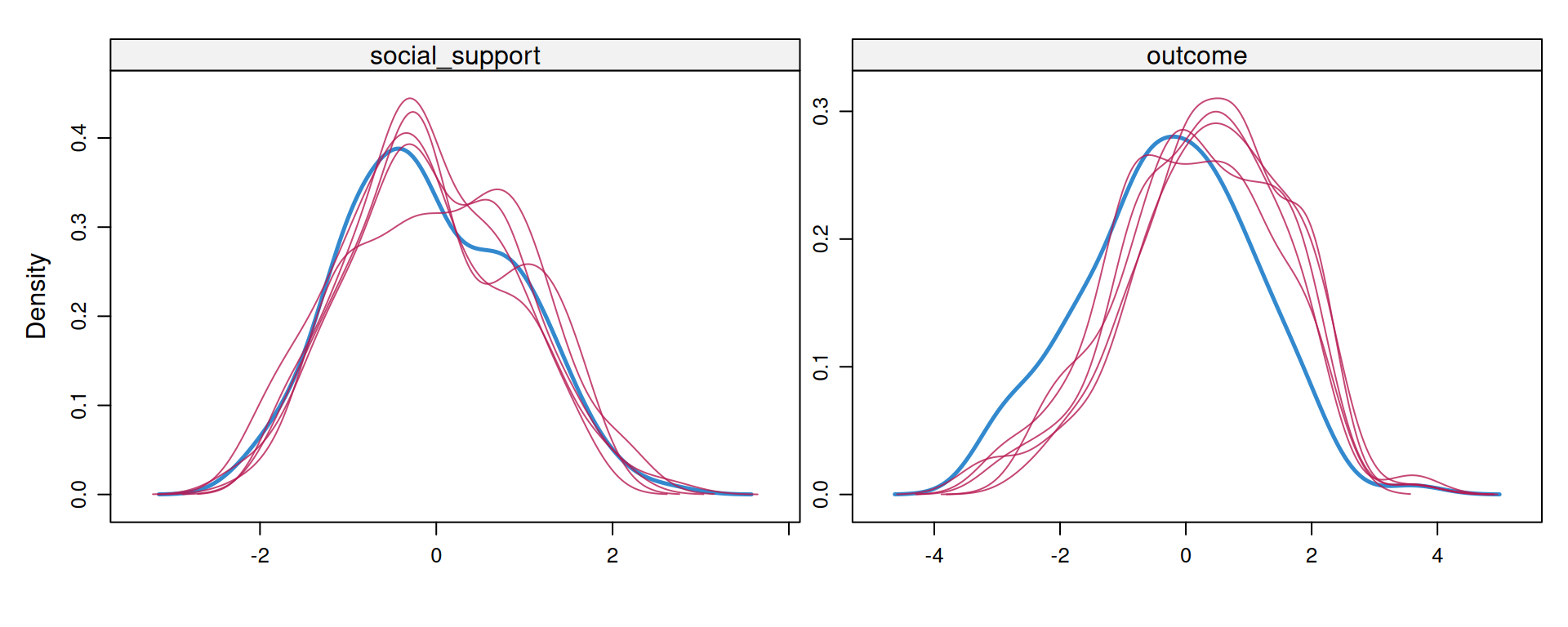
Blue = observed data distribution. Red lines = imputed distributions (one per imputation). The shapes should be plausible and similar to the observed data, but not necessarily identical.
Step 4: Analyze Each Imputed Dataset
This fits the same regression model on all 5 imputed datasets. Each produces slightly different estimates because the imputed values differ.
Step 5: Pool with Rubin’s Rules
| term | estimate | std.error | 2.5 % | 97.5 % | p.value |
|---|---|---|---|---|---|
| (Intercept) | -0.088 | 0.199 | -0.537 | 0.362 | 0.670 |
| treatment | -0.522 | 0.168 | -0.861 | -0.183 | 0.003 |
| age | -0.269 | 0.090 | -0.453 | -0.084 | 0.006 |
| female | 0.598 | 0.168 | 0.262 | 0.935 | 0.001 |
| baseline_severity | 0.372 | 0.091 | 0.187 | 0.556 | 0.000 |
| social_support | -0.392 | 0.093 | -0.582 | -0.202 | 0.000 |
The pooled standard errors incorporate both sampling variability (within) and imputation uncertainty (between).
Complete-Case vs. MI
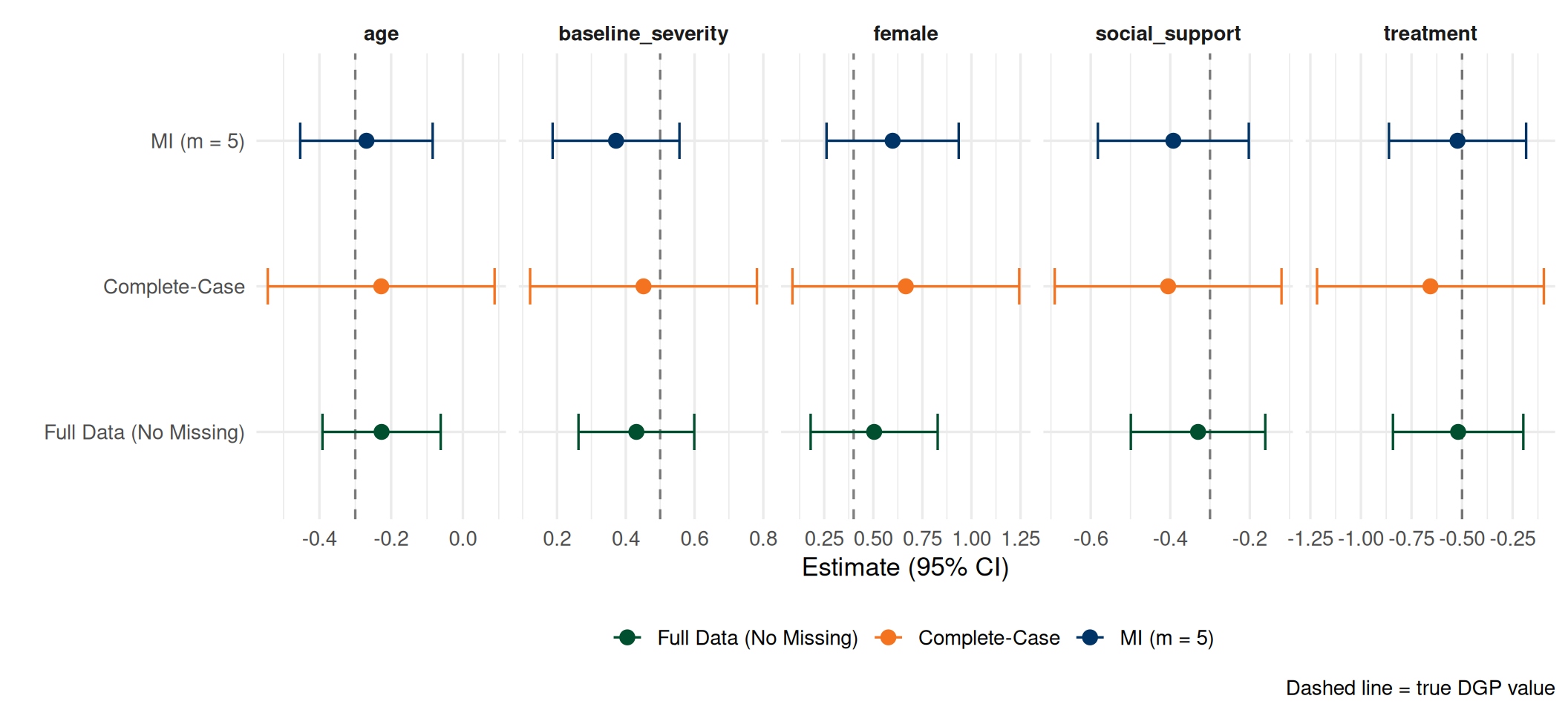
- MI recovers estimates close to the full-data truth
- MI standard errors properly account for imputation uncertainty (between)
- Complete-case discards all partially observed rows, losing efficiency
MI: Practical Guidelines
How many imputations (m)?
- Old rule of thumb: m = 5
- Other recommendations: m ≥ 20 [@White_2010]
- Depends on the fraction of missing information
Include auxiliary variables:
- Variables that predict missingness
- Variables that correlate with the incomplete variable
- Even if they are not in your analysis model
Always check:
- Convergence - trace plots should show no trends
- Plausibility - imputed distributions should be reasonable
- Sensitivity - try different imputation models
Report in your paper:
- Number of imputations (m)
- Imputation method (e.g., PMM, logistic)
- Variables included in the imputation model
- Software and version used
The Imputation Model Should Be at Least as General as the Analysis Model
If your analysis includes interactions or nonlinear terms, include them in the imputation model too.
Other MI Packages in R
Beyond mice, several R packages offer alternative approaches to multiple imputation:
| Package | Approach | Best For | Link |
|---|---|---|---|
| mice | Chained equations (FCS) | General-purpose, flexible variable-by-variable imputation | CRAN |
| Amelia | Joint multivariate normal (EMB algorithm) | Large datasets, continuous/normal data, time-series cross-sectional | CRAN |
| jomo | Joint modelling with random effects | Multilevel / clustered data, mixed variable types | CRAN |
| missForest | Random forest (single imputation) | Nonlinear relationships, mixed types, no distributional assumptions | CRAN |
| miceadds | Extensions to mice | Multilevel imputation, two-level models, plausible values | CRAN |
| mi | Bayesian regression | Diagnostics-rich workflow, automatic transformations | CRAN |
How to choose?
- Flat data, mixed types \(\rightarrow\)
mice(chained equations, most flexible) - Mostly continuous, large N \(\rightarrow\)
Amelia(fast joint model, assumes MVN) - Multilevel / clustered \(\rightarrow\)
jomoormiceadds(proper random-effects imputation) - Complex nonlinear patterns \(\rightarrow\)
missForest(tree-based, but single imputation only)
FIML with lavaan
Full Information Maximum Likelihood (FIML)
FIML estimates model parameters by maximizing the likelihood using all available data, even from incomplete cases.
Unlike MI, FIML does not fill in missing values. Instead, it computes the likelihood for each case based on whatever variables are observed for that case.
Key properties:
- Produces unbiased estimates under MAR
- Uses all observed information, no cases discarded
- Handled natively in lavaan or Mplus with a single argument
- No need to create or manage multiple datasets
- Equivalent to MI asymptotically (same results, different path)
A 5-Item Anxiety Scale
We simulate a one-factor CFA with 5 items measuring latent anxiety:
library(lavaan)
pop_model <- "
anxiety =~ 0.80*item1 + 0.70*item2 + 0.60*item3 + 0.75*item4 + 0.65*item5
anxiety ~~ 1*anxiety
item1 ~~ 0.36*item1
item2 ~~ 0.51*item2
item3 ~~ 0.64*item3
item4 ~~ 0.4375*item4
item5 ~~ 0.5775*item5
"
set.seed(2026)
fiml_complete <- simulateData(pop_model, sample.nobs = 2000)True standardized loadings: \(\lambda_1 = .80\), \(\lambda_2 = .70\), \(\lambda_3 = .60\), \(\lambda_4 = .75\), \(\lambda_5 = .65\)
Survey Fatigue Strikes (MAR)
Participants who score higher on item 1 (more anxious) are more likely to skip later items, a realistic survey fatigue pattern:
fiml_data <- fiml_complete |>
as_tibble() |>
mutate(
# Later items have progressively more missingness
item3 = ifelse(rbinom(n(), 1, plogis(-2.0 + 0.8 * item1)) == 1, NA, item3),
item4 = ifelse(rbinom(n(), 1, plogis(-1.5 + 0.8 * item1)) == 1, NA, item4),
item5 = ifelse(rbinom(n(), 1, plogis(-1.0 + 0.8 * item1)) == 1, NA, item5)
)This is MAR: missingness depends on item1 (always observed), not on the missing items themselves.
| Item | N_Missing | Pct |
|---|---|---|
| item3 | 277 | 13.9 |
| item4 | 440 | 22.0 |
| item5 | 577 | 28.8 |
The CFA Model in lavaan
The model specification is identical for all three analyses. Only the missing argument changes:
cfa_model <- "
anxiety =~ item1 + item2 + item3 + item4 + item5
"
# 1. Full data (truth <U+2014> no missing values)
fit_true <- cfa(cfa_model, data = fiml_complete)
# 2. Listwise deletion (default)
fit_lw <- cfa(cfa_model, data = fiml_data, missing = "listwise")
# 3. FIML <U+2014> just add one argument
fit_fiml <- cfa(cfa_model, data = fiml_data, missing = "fiml")That’s It
The only difference is missing = "fiml". lavaan handles everything internally: no imputed datasets, no pooling, no extra steps.
Listwise vs FIML: Factor Loadings
| Item | DGP | Full Data | Listwise | FIML | LW (SE) | FIML (SE) |
|---|---|---|---|---|---|---|
| item1 | 0.80 | 0.794 | 0.781 | 0.798 | 0.018 | 0.013 |
| item2 | 0.70 | 0.661 | 0.645 | 0.667 | 0.022 | 0.016 |
| item3 | 0.60 | 0.568 | 0.568 | 0.565 | 0.025 | 0.020 |
| item4 | 0.75 | 0.748 | 0.757 | 0.754 | 0.018 | 0.015 |
| item5 | 0.65 | 0.665 | 0.632 | 0.650 | 0.023 | 0.019 |
| Full Data | Listwise | FIML | |
|---|---|---|---|
| N used | 2000 | 1029 | 2000 |
Listwise discards every case with any missing item. FIML uses all 2000 cases.
Listwise vs FIML: Visual Comparison
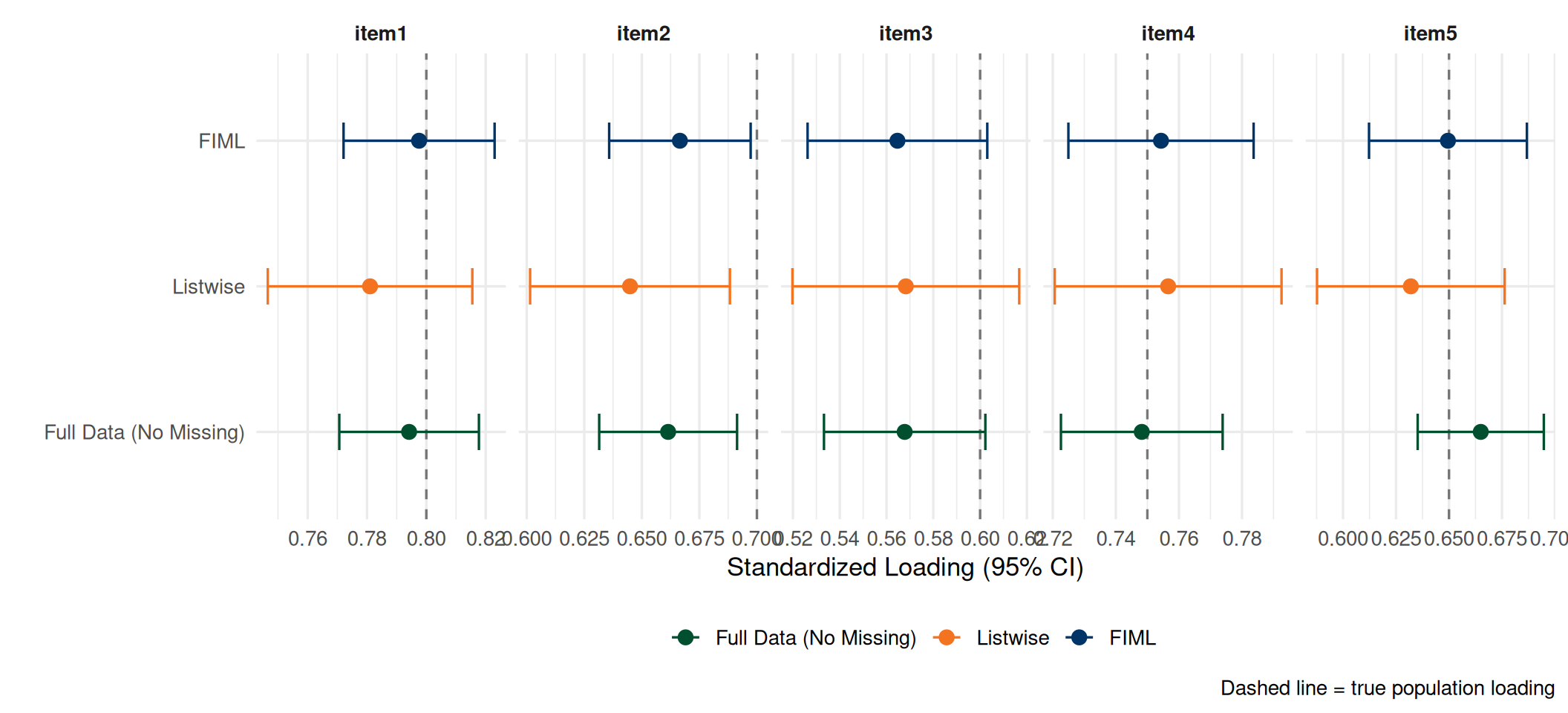
- FIML recovers loadings close to the truth — especially for items 3–5 where missingness was highest
- Listwise confidence intervals are wider (less efficient) due to reduced sample size
Model Fit: Listwise vs FIML
| Index | Full Data | Listwise | FIML |
|---|---|---|---|
| <U+03C7><U+00B2> | 6.720 | 1.704 | 1.930 |
| df | 5.000 | 5.000 | 5.000 |
| p-value | 0.242 | 0.888 | 0.859 |
| CFI | 0.999 | 1.000 | 1.000 |
| TLI | 0.999 | 1.004 | 1.002 |
| RMSEA | 0.013 | 0.000 | 0.000 |
| SRMR | 0.008 | 0.006 | 0.004 |
FIML does not improve model fit magically — it provides more accurate estimates of the population parameters by using all available information.
lavaan 0.6-21 ended normally after 21 iterations
Estimator ML
Optimization method NLMINB
Number of model parameters 15
Number of observations 2000
Number of missing patterns 8
Model Test User Model:
Test statistic 1.930
Degrees of freedom 5
P-value (Chi-square) 0.859
Model Test Baseline Model:
Test statistic 2494.629
Degrees of freedom 10
P-value 0.000
User Model versus Baseline Model:
Comparative Fit Index (CFI) 1.000
Tucker-Lewis Index (TLI) 1.002
Robust Comparative Fit Index (CFI) 1.000
Robust Tucker-Lewis Index (TLI) 1.003
Loglikelihood and Information Criteria:
Loglikelihood user model (H0) -11008.303
Loglikelihood unrestricted model (H1) -11007.338
Akaike (AIC) 22046.607
Bayesian (BIC) 22130.620
Sample-size adjusted Bayesian (SABIC) 22082.965
Root Mean Square Error of Approximation:
RMSEA 0.000
90 Percent confidence interval - lower 0.000
90 Percent confidence interval - upper 0.017
P-value H_0: RMSEA <= 0.050 1.000
P-value H_0: RMSEA >= 0.080 0.000
Robust RMSEA 0.000
90 Percent confidence interval - lower 0.000
90 Percent confidence interval - upper 0.019
P-value H_0: Robust RMSEA <= 0.050 1.000
P-value H_0: Robust RMSEA >= 0.080 0.000
Standardized Root Mean Square Residual:
SRMR 0.004
Parameter Estimates:
Standard errors Standard
Information Observed
Observed information based on Hessian
Latent Variables:
Estimate Std.Err z-value P(>|z|) Std.lv Std.all
anxiety =~
item1 1.000 0.780 0.798
item2 0.851 0.032 27.009 0.000 0.664 0.667
item3 0.713 0.033 21.486 0.000 0.556 0.565
item4 0.984 0.035 28.188 0.000 0.767 0.754
item5 0.827 0.036 22.899 0.000 0.645 0.650
Intercepts:
Estimate Std.Err z-value P(>|z|) Std.lv Std.all
.item1 -0.005 0.022 -0.245 0.807 -0.005 -0.005
.item2 0.007 0.022 0.313 0.754 0.007 0.007
.item3 0.010 0.023 0.441 0.659 0.010 0.010
.item4 -0.004 0.025 -0.146 0.884 -0.004 -0.004
.item5 0.002 0.025 0.082 0.935 0.002 0.002
Variances:
Estimate Std.Err z-value P(>|z|) Std.lv Std.all
.item1 0.348 0.019 18.352 0.000 0.348 0.364
.item2 0.551 0.022 25.595 0.000 0.551 0.556
.item3 0.661 0.025 26.203 0.000 0.661 0.681
.item4 0.446 0.023 19.698 0.000 0.446 0.431
.item5 0.570 0.025 22.381 0.000 0.570 0.578
anxiety 0.608 0.032 18.936 0.000 1.000 1.000FIML vs MI: When is common to use (not a rule)?
FIML:
- Your model is a CFA, SEM, or path model
- You’re already using lavaan / Mplus
- You want a simple, single-step solution
- You prefer not to manage multiple datasets
MI:
- Your analysis is not SEM-based (e.g., regression, ANOVA, mixed models)
- You need to use software that doesn’t support FIML
- You want to include many auxiliary variables flexibly
- You need to do sensitivity analyses with different imputation models
Important
Both MI and FIML assume MAR. If you suspect MNAR, you need sensitivity analysis, neither method alone can save you.
Wrapping Up: Your Complete Toolkit
| Method | When to Use | Assumption | Bias? |
|---|---|---|---|
| Complete-case | Quick look, MCAR only | MCAR | No (under MCAR) |
| MI (MICE) | Any model, any software | MAR | No (under MAR) |
| FIML | CFA / SEM models | MAR | No (under MAR) |
| Sensitivity | When MNAR is suspected | MNAR | Depends |
Your workflow:
- Diagnose Why is data missing? (MCAR / MAR / MNAR)
- Choose Pick the right tool for your model and mechanism
- Check Convergence, plausibility, sensitivity
- Report Be transparent about what you did and why
References
- Little, R. J. A., & Rubin, D. B. (2002). Statistical Analysis with Missing Data (2nd ed.). Wiley.
- Rubin, D. B. (1976). Inference and missing data. Biometrika, 63(3), 581–592.
- Enders, C. K. (2010). Applied Missing Data Analysis. Guilford Press.
- Schafer, J. L., & Graham, J. W. (2002). Missing data: Our view of the state of the art. Psychological Methods, 7(2), 147–177.
- van Buuren, S., & Groothuis-Oudshoorn, K. (2011). mice: Multivariate Imputation by Chained Equations in R. Journal of Statistical Software, 45(3), 1–67.
- Rosseel, Y. (2012). lavaan: An R package for structural equation modeling. Journal of Statistical Software, 48(2), 1–36.
- White, I. R., Royston, P., & Wood, A. M. (2011). Multiple imputation using chained equations: Issues and guidance for practice. Statistics in Medicine, 30(4), 377–399.
Questions?

EPH 752 Advanced Research Methods